library(tidymodels)── Attaching packages ────────────────────────────────────── tidymodels 1.4.1 ──✔ broom 1.0.12 ✔ recipes 1.3.1
✔ dials 1.4.2 ✔ rsample 1.3.2
✔ dplyr 1.2.0 ✔ tailor 0.1.0
✔ ggplot2 4.0.1 ✔ tidyr 1.3.2
✔ infer 1.1.0 ✔ tune 2.0.1
✔ modeldata 1.5.1 ✔ workflows 1.3.0
✔ parsnip 1.4.1 ✔ workflowsets 1.1.1
✔ purrr 1.2.1 ✔ yardstick 1.3.2 ── Conflicts ───────────────────────────────────────── tidymodels_conflicts() ──
✖ purrr::discard() masks scales::discard()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
✖ recipes::step() masks stats::step()library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ lubridate 1.9.5 ✔ tibble 3.3.1
✔ readr 2.1.6 ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ readr::col_factor() masks scales::col_factor()
✖ purrr::discard() masks scales::discard()
✖ dplyr::filter() masks stats::filter()
✖ stringr::fixed() masks recipes::fixed()
✖ dplyr::lag() masks stats::lag()
✖ readr::spec() masks yardstick::spec()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(ggplot2)
rngseed <- 1234
set.seed(rngseed)
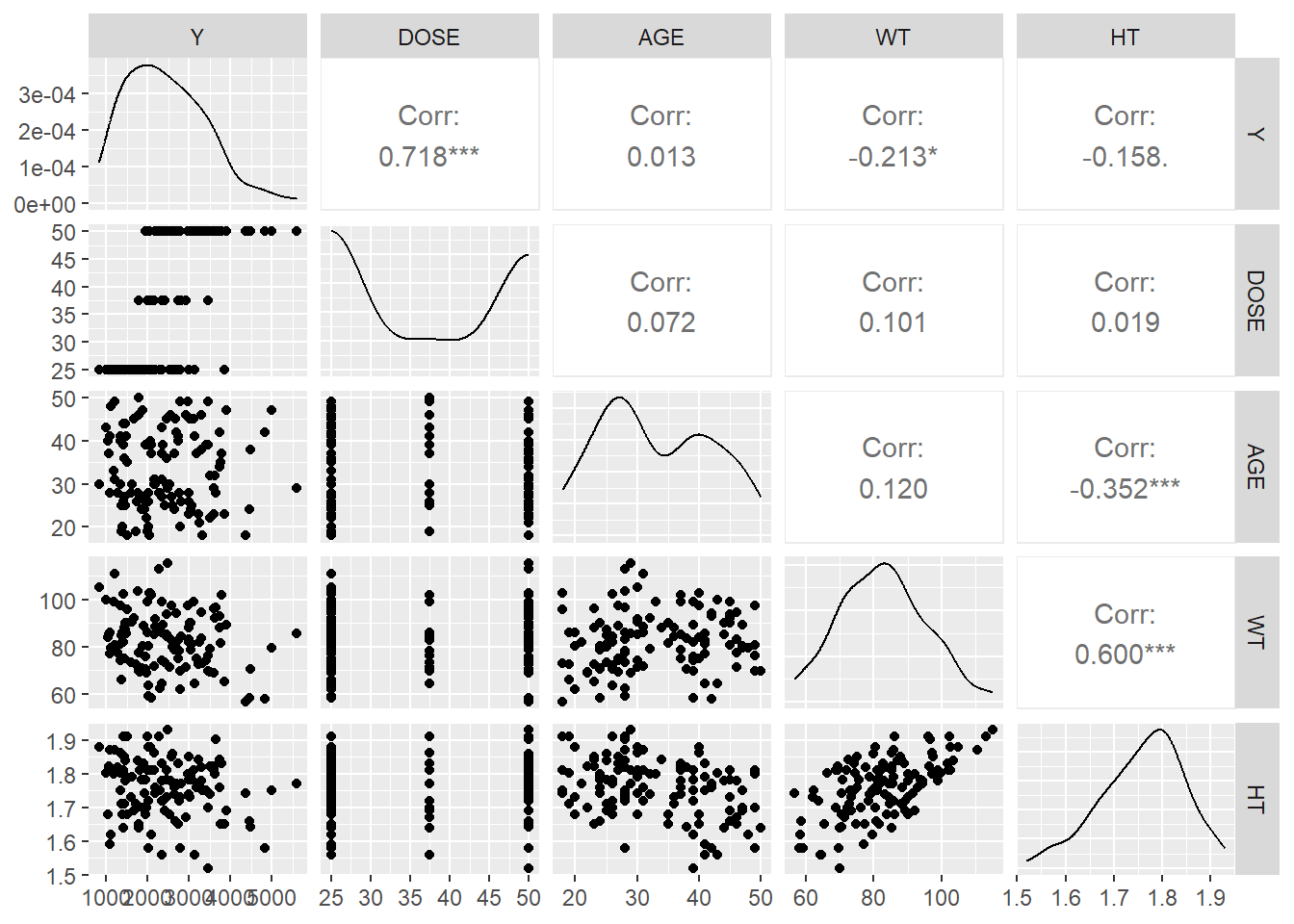
dat <- readRDS("dat_clean.rds")
glimpse(dat)Rows: 120
Columns: 7
$ Y <dbl> 2690.52, 2638.81, 2149.61, 1788.89, 3126.37, 2336.89, 3007.20, 27…
$ DOSE <dbl> 25.0, 25.0, 25.0, 25.0, 25.0, 25.0, 25.0, 25.0, 25.0, 25.0, 25.0,…
$ AGE <dbl> 42, 24, 31, 46, 41, 27, 23, 20, 23, 28, 46, 22, 43, 50, 19, 26, 3…
$ SEX <fct> 1, 1, 1, 2, 2, 1, 1, 1, 1, 1, 1, 1, 2, 2, 1, 1, 1, 1, 1, 1, 1, 1,…
$ RACE <fct> 2, 2, 1, 1, 2, 2, 1, 88, 2, 1, 1, 1, 1, 1, 2, 2, 1, 1, 1, 1, 1, 1…
$ WT <dbl> 94.3, 80.4, 71.8, 77.4, 64.3, 74.1, 87.9, 61.9, 65.3, 103.5, 83.0…
$ HT <dbl> 1.769997, 1.759850, 1.809847, 1.649993, 1.560052, 1.829862, 1.850…